An optimized protocol for rapid, sensitive and robust on-bead ChIP-seq from primary cells: STAR Protocols

Analysis of the Chromosomal Localization of Yeast SMC Complexes by Chromatin Immunoprecipitation | SpringerLink
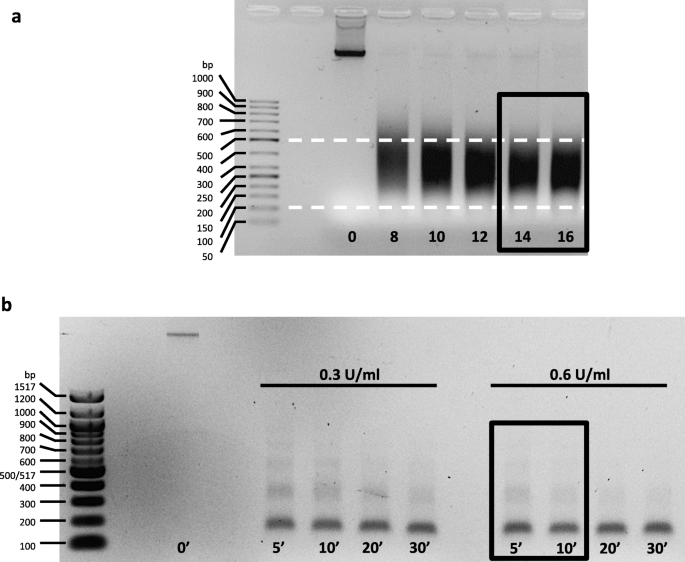
ChIP-seq and RNA-seq for complex and low-abundance tree buds reveal chromatin and expression co-dynamics during sweet cherry bud dormancy | SpringerLink